Oligo Synthesis FAQ
Oligo Synthesis FAQ Catalog:
- What does OD value mean?
- How is synthesized DNA / RNA quantified?
- How is the OD value of synthesized DNA / RNA determined and calculated?
- How are synthesized DNA dry powders diluted and preserved?
- How are synthesized RNA dry powders diluted and preserved?
- How do I dissolve Oligo DNA/RNA smaller than 15 mer?
- How are synthesized Oligo products electrophoresed?
- How are synthesized fluorescently labeled probes stored?
- Will a synthesized DNA solution degrade if left at room temperature for a few days?
- Oligo solution has OD260 / OD280 < 1.8, does this mean the solution is impure?
- What are the purification levels of Oligo products? What is the difference?
- How do you calculate the molarity of a synthesized Oligo DNA product?
- The synthesized Oligo was dissolved and found to have a small amount of precipitate, will this affect subsequent experiments?
- When Oligo DNA of the same length is subjected to PAGE electrophoresis, why are the positions of the bands inconsistent?
- Is there a phosphate group at the 5′ and 3′ ends of synthetic Oligo DNA?
- When performing antisense nucleic acid experiments, do you need to modify all of the DNA strands with S substitutions, and are there other ways to increase the stability of nucleic acids besides S substitutions?
- What principles should be followed in the design of dual-labeled fluorescent probes?
- How should quenching groups and fluorescent groups be paired when modifying fluorescent probes?
- How should fluorescent group modifications be selected for multiprobe assays?
- How to choose the quenching group of TAMRA and BHQ?
- What is the difference between FITC, 6-FAM, and 5-FAM markers?
- The PCR product was cloned and sequenced and found to have an error in a base at the primer.
- Why are flat PCR products harder to clone?
- What is the length of DNA / RNA that can be synthesized by Aceri?
What does OD value mean?
OD is an abbreviation for optical density, which indicates the density of light absorbed by the detected object.
How is synthesized DNA / RNA quantified?
Because nucleic acids absorb strongly near 260 nm, they can be quantified by UV spectrophotometry based on this property and expressed as an OD value.1 OD means that if a solution of DNA/RNA is placed in a cuvette with an optical path length of 1 cm, and if its absorbance at 260 nm is 1, then the amount of DNA/RNA dissolved in 1 ml of the solution is said to be 1 OD. The mass of synthesized DNA/RNA for one OD is approximately 33 μg (the unit of total amount, not concentration). The mass of synthesized DNA/RNA for 1 OD is approximately 33 μg (OD is a unit of total amount, not concentration).
How is the OD value of synthesized DNA / RNA determined and calculated?
The DNA/RNA solution is diluted appropriately so that the absorbance measurement is within the precise range allowed by the instrument, and then the total OD value of the solution is calculated based on the total volume of the original solution and the number of times it was diluted. For example, if a 300 μl DNA/RNA solution is diluted 5-fold and the measured value is 1.0, the total OD value of the solution is: 0.3 ml × 5 × 1.0 OD / ml = 1.5 OD (if the measured value is greater than 1.5, it is recommended to re-dilute the solution before measurement in order to ensure the accuracy of the measurement).
How are synthesized DNA dry powders diluted and preserved?
For long term storage, it is recommended to store the dry powder directly or dissolve it in sterile TE buffer (10 mM Tris pH 8.0, 0.1 M EDTA) to a higher concentration (e.g., 100 µM) and store at -20°C or -80°C, avoiding repeated freezing and thawing (do not store in a refrigerator with an automatic defrost function). It can also be diluted in sterile, enzyme-free water or to a working concentration (10 µM), but the effective shelf-life will be shortened.
How are synthesized RNA dry powders diluted and preserved?
The dry powder is recommended to be stored at -80°C, if not, please store at -20°C. Usually it can be stored for 2 years. RNA in solution state is best stored at -80°C, if not, store at -20°C.
How do I dissolve Oligo DNA/RNA smaller than 15 mer?
Oligo DNA / RNA length ≤15 mer or sequence-specific cases will have reduced solubility (Milli-Q water may be insoluble) and TE (pH 7.5 - 8.0) is recommended for solubilization.
How are synthesized Oligos products electrophoresed?
Single-stranded Oligo tends to form complex three-dimensional structures and is therefore not suitable for agarose electrophoresis (multiple bands may appear) and should not be stained with EB, GoldView or CelRed. Electrophoresis should be performed using 15% Denaturing PAGE Gel, and visualized by imaging on a silica gel fluorescent plate with a UV lamp (254 nm).
How are synthesized fluorescently labeled probes stored?
Fluorescent probes should be stored under strict light protection; dry powder can be stored at -20℃ for 2 years without degradation, but care should be taken to avoid repeated freezing and thawing.
Will a synthesized DNA solution degrade if left at room temperature for a few days?
In general, a 100 µM concentration of Oligo DNA solution at room temperature (25°C - 30°C) for 3-4 days has little effect, but it is best not to leave it for more than a week. The exact amount of time depends on the degree of sterilization of the synthesis vessels and reagents.
Oligo solution has OD260 / OD280 < 1.8, does this mean the solution is impure?
Nucleic acids have strong absorption near 260 nm, and proteins have strong absorption near 280 nm. When the proportions of A, G, C, and T in the sequences are roughly the same, the OD260 / OD280 ratio can be used to evaluate whether there is any protein residue in the nucleic acids (the ratio is in the range of 1.8 to 2.2), such as nucleic acids extracted from organisms. However, the synthesized Oligo sequence is very short, in which A, G, C, and T bases account for different proportions, and since the molar extinction coefficients of the bases are different, the OD260/OD280 ratios of Oligo sequences composed of different bases are also different. For example, when the sequence has a high content of C and T bases, the ratio is much lower than 1.8. In addition, the order of the bases in the sequence also affects the ratio. Therefore, it is not possible to evaluate the purity of synthesized Oligo based on the OD260 / OD280 ratio.
| base percentage | OD260 / OD280 |
| 100% A Base | 2.5 |
| 100% G Base | 1.85 |
| 100% C Base | 1.15 |
| 100% T Base | 1.14 |
| A, G, C, T (25% each) | 1.66 |
What are the purification levels of Oligo products? What is the difference?
I. Desalting grade: Simple desalting to remove salts and ions generated during synthesis, suitable for shorter sequences with less demanding downstream experiments.
Second, OPC purification grade: the small column of C18 resin is used for purification, which can remove the short-stranded DNA that fails to be synthesized.
PAGE level: According to the charge and molecular size, the target sequence can be distinguished from the failed longer sequences, and within a certain range, it can effectively remove the N-1 impurity chain.
HPLC grade: It can separate N-1 impurities more effectively through the combined purification of ion exchange column and reverse C18 column.
| Purification method | appliance | Range of purified bases (bases) | fineness |
| desalination | Sequencing, conventional PCR, hybridization DNA microarrays, SNP analysis, etc. | 10 - 60 | Related to sequence length |
| OPC level | AFLP, OLA, diagnostic PCR, antisense nucleic acids, traditional modifications, etc. | 2 - 60 | 80% Above |
| PAGE Level | Primers for molecular diagnostics, quantitative PCR, cloning, subcloning PCR, gene synthesis, gel migration evaluation, etc. | 10 or more | 90% - 95% |
| HPLC grade | In situ hybridization, miRNA, siRNA, quantitative PCR, capillary sequencing, primers for molecular diagnostics, etc. | 2 - 60 | 95% Above |
| HPLC + PAGE grade | X-Ray Crystallographic studies, NMR, 3D structural studies, difficult cloning and gene synthesis, etc. | 10 or more | Highest purity |
How do you calculate the molarity of a synthesized Oligo DNA product?
Since each base has a different light-absorbing capacity (molar extinction coefficient ε260), a more accurate formula for calculating the Oligo DNA nmol number is as follows:
nmol number = total solution OD * 106 / ε260
ε260 = (15.2 * number of A bases) + (11.5 * number of G bases) + (7.4 * number of C bases) + (8.3 * number of T bases)
The synthesized Oligo was dissolved and found to have a small amount of precipitate, will this affect subsequent experiments?
Occasionally, trace amounts of C18 resin may enter the product during OPC purification. The resin does not affect any of the reaction results and can be centrifuged slightly and the supernatant used.
When Oligo DNA of the same length is subjected to PAGE electrophoresis, why are the positions of the bands inconsistent?
Different base compositions result in inconsistent electrophoretic speeds for Oligo DNA; different DNA stereo structures also have different electrophoretic speeds. (Usually occurs in shorter sequences, with less variation among long-stranded Oligo DNA).
Is there a phosphate group at the 5' and 3' ends of synthetic Oligo DNA?
If not required, the synthesized Oligo DNA is normally -OH without a phosphate group at the end. If an additional phosphate group is required, it can be specified when placing the order, in which case the cost of phosphorylation (PO4 modification) will be charged.
When performing antisense nucleic acid experiments, is it necessary to modify all of the DNA strand with the S-substitution, and is there any other way to increase the stability of nucleic acids besides the S-substitution?
S-substitution modification increases the stability of the nucleic acid in the cell and prevents degradation by nuclease. However, complete modification of the entire strand is not recommended (except for injection into live animals), as this will reduce the Tm value and the binding efficiency of the antisense DNA to the target sequence. Phosphorothioated Bonds can be inserted at both ends of the DNA fragments to increase both the stability of the DNA and the ability of the antisense DNA to bind to the target sequence.
In addition to S-generation modifications, 2' - OMe - RNA can also be used to increase stability by forming a chimeric antisense nucleic acid with S-generation DNA, which both enhances the stability of the antisense nucleic acid and increases the affinity for the target sequence (the affinity of 2' - OMe - RNA for RNA is much greater than that of DNA for RNA). In addition, 5 - Me - dC can also be added to S-generation antisense nucleic acids due to its ability to enhance the stability of the DNA double helix.
What principles should be followed in the design of dual-labeled fluorescent probes?
● The probe should be located between the two primers and can be either a forward or reverse probe;
● The content of bases G and C in the probe should preferably be controlled between 40% and 60%; avoid strings of the same base, especially base G;
● Base G should not appear at the 5' end; the Tm value of the probe should be 8-10℃ higher than that of the primer, preferably between 68 and 70℃;
● When the length of the probe exceeds 30 bases, it is recommended to put the quenching group in the middle to prevent the fluorescence background from being too high, and then the 3' end of the probe should be blocked with a phosphoryl group to prevent the probe from extending during the PCR reaction.
How should quenching groups and fluorescent groups be paired when modifying fluorescent probes?
The role of the quenching group is to quench the fluorescence of the reporter group, so the fluorescence spectrum of the quenching group should be similar to the absorption spectrum of the reporter group. Reference can be made to the following figure:
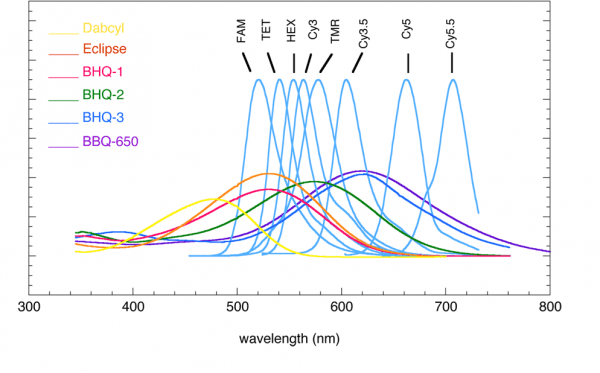

How should fluorescent group modifications be selected for multiprobe assays?
The traditional fluorescence quantitative PCR instrument is usually four fluorescence detection channels, each channel detects different fluorescence signals, can refer to the following table to match:
| excitation wavelength | Detection wavelength | fluorescent marker | |
| Channel 1 | 450 - 490 nm | 515 - 530 nm | FAM, SYBR Green |
| second channel | 500 - 535 nm | 560 - 580 nm | vic, hex, toe, tamra, tet, cy3 |
| third channel | 555 - 585 nm | 610 - 650 nm | ROX, Texas-Red |
| Corridor 4 | 620 - 650 nm | 675 - 730 nm | Cy5 |
How to choose the quenching group of TAMRA and BHQ?
TAMRA is a fluorescent dye and emits its own fluorescence at higher wavelengths, so the probe fluorescence background is relatively high. The BHQ series are non-fluorescent dyes that only quench the reporter group and do not emit fluorescence themselves, resulting in a low probe fluorescence background, a higher signal-to-noise ratio, and higher detection sensitivity.
TAMRA has a narrow absorption spectrum with fewer types of quenching groups, whereas the BHQ series of dyes have a wider absorption spectrum with more types of quenching groups. Therefore, the BHQ series of dyes are commonly used for quenching moieties in multiplex PCR assays.
What is the difference between FITC, 6-FAM, and 5-FAM markers?
All three are fluorescein-labeled (Fluorescein), and the chromophore is fluorescein; however, structurally, 5 - FAM and 6 - FAM are isomers of each other; FITC is linked to Oligo by a thiourea bond), whereas the former is an amide bond, and there is no difference in usual use.
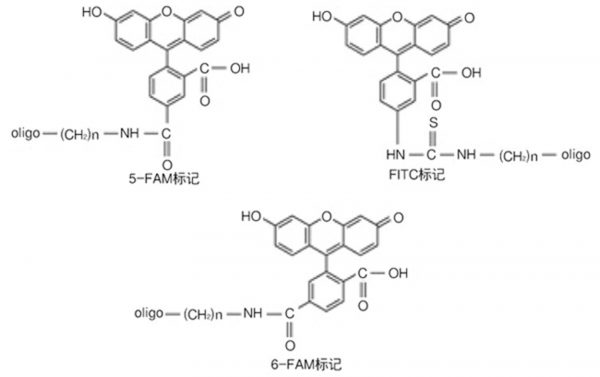

The PCR product was cloned and sequenced and found to have an error in a base at the primer.
Base errors at primers can be troubleshooted in several ways:
● Artificially entering the sequence incorrectly (synthetic report card can be checked);
● Whether the purification level is suitable for downstream experiments;
● Reselect clones for sequencing (primer purity is unlikely to be 100%; it is possible that clones with PCR products amplified by impurity primers were selected).
If there is still a problem through the above troubleshooting, Acres will re-synthesize the primers free of charge.
Why are flat PCR products harder to clone?
Typical PCR primers do not have a phosphate group at the 5′ end, and therefore the amplified PCR product does not have a phosphate group at the 5′ end. When cloned into dephosphorylated end-smoothing vectors, they cannot be cloned in, and when cloned into non-dephosphorylated end-smoothing vectors, the background is extremely high. This can be achieved by phosphorylating (PO4-modifying) the 5′ end of the PCR product.
What is the length of DNA / RNA that can be synthesized by Aceri?
The length of synthesized DNA ranges from 1 to 200 bases; the length of synthesized RNA ranges from 1 to 120 bases. All products use TOF-MS for molecular weight determination to ensure the accuracy of synthesis (when synthesized DNA sequences are long, it is difficult to ensure that each base is correct due to the limitations of synthesis and purification methods).
