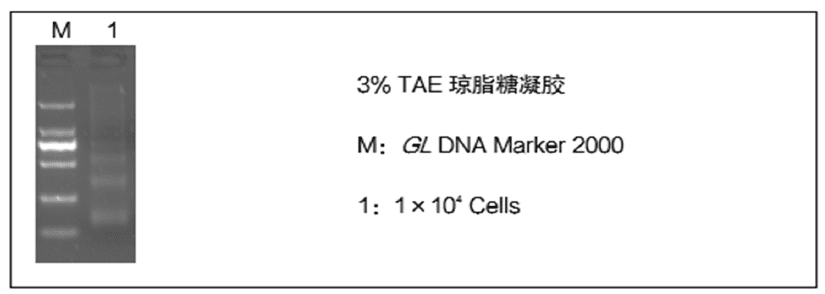
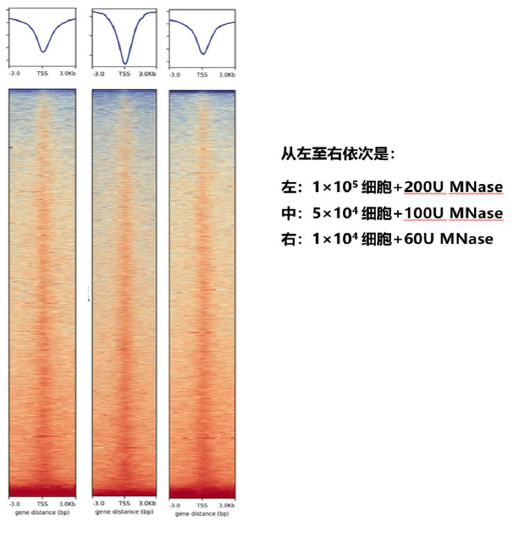
This product is a kit developed based on Micrococcal Nuclease (MNase), which is suitable for nucleosome localization and occupancy analysis, research on gene transcription and regulatory mechanisms, epigenetic and chromatin remodeling studies, exploration of disease mechanisms, and evolutionary and developmental biology applications. The starting volume of this product can be as low as 104~105Individual cells.
It utilizes the properties of micrococcal nuclease to selectively digest naked DNA and retain nucleosome-encapsulated DNA fragments for subsequent library construction and high-throughput sequencing analysis.
The reaction system of this product has been carefully optimized, all reagents should be used as provided in this product, it is not recommended to change any of the reaction components or use other equivalent products to replace the components in this product, in order to avoid obtaining bad experimental results. If substitution is required, please verify first.
1. High specificity: MNase Enzyme accurately digests naked DNA, preserving intact nucleosome DNA fragments (~150 bp sized monomer peaks are prominent).
2. Dual enzyme synergistic digestion: MNase Enzyme + Exonuclease III Enzyme is used in combination to reduce background noise and improve the resolution of nucleosome localization.
3. Optimization of the whole process: premixed buffer ensures the digestion efficiency, and DNA Clean Beads purification replaces phenol-chlorine extraction, which is safer to operate and has a higher purification recovery rate.
Package 2-1 is grouped as follows (stored at 4°C):
| individual parts making up a compound | AG12572 | AG12573 |
| DNA Clean Beads | 540 μl | 1.7 ml |
Package 2-2 is grouped as follows (stored at -20°C):
| individual parts making up a compound | AG12572 | AG12573 |
| RNase A ( 10 mg / ml ) | 6 μl | 20 μl |
| Proteinase K ( 20 mg / ml ) | 20 μl | 60 μl |
| Lysis Buffer | 200 μl | 600 μl |
| Wash Buffer Ⅰ | 1 ml x 2 pcs | 1.5 ml x 4 pcs |
| 1% Digitonin | 4 μl | 12 μl |
| Dilution Buffer | 300 μl | 1 ml |
| MNase Enzyme | 4 μl | 12 μl |
| Exonuclease III Enzyme ( 5 U/μl ) | 4 μl | 12 μl |
| MNase Digestion Buffer | 200 μl | 600 μl |
| MNase Stop Buffer | 24 μl | 75 μl |
| 10% SDS | 24 μl | 75 μl |
| Nuclease free water | 1 ml | 1 ml x 2 pcs |
Note: Recommended use AccuNext DNA Library Preparation Kit (Illumina) (Code No. AG12535 / AG12536) is available for subsequent library construction, and can be purchased with the AccuNext CDI Junction Primers (Illumina, for DNA libraries) (Code No. AG12529 / AG12530 / AG12531),AccuNext Short junction primers (Illumina, for DNA libraries) (Code No. AG12537 / AG12538).
Package 2-2 -20°C Storage
Transportation Temperature: Package 2-1 Ice Pack Transportation
Package 2-2 Dry Ice Transportation or -20℃ Ice Pack Transportation
Stability: Unopened reagents are valid for 12 months, after opening, they are stored according to the requirements of the components (e.g. MNase Enzyme is stored at -20℃ for ≤6 months after dispensing).