Product Description
This product is a kit for obtaining the unknown sequences of the flanks of genomic DNA based on the known sequences on the genomic DNA, which is improved on the principle of TAIL-PCR (Thermal Asymmetric Interlaced PCR). This method is characterized by high efficiency, high specificity, high sensitivity and easy operation compared with other traditional chromosome stepping methods (e.g. reverse PCR, junction method, etc.).
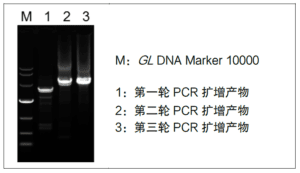
The flanking sequences of the known sequences were obtained after three rounds of PCR amplification using the four concatenated primers provided in the kit, i.e., TP Primer 1 (TP1), TP Primer 2 (TP2), TP Primer 3 (TP3), and TP Primer 4 (TP4), respectively, and paired with the specific primers designed according to the known sequences, i.e., Specific Primer 1 (SP1), Specific Primer 2 (SP2), and Specific Primer 3 (SP3) for thermal asymmetric PCR amplification. Specific Primer 1 (SP1), Specific Primer 2 (SP2) and Specific Primer 3 (SP3) were used for thermal asymmetric PCR amplification, and the flanking sequences of the known sequences could be obtained after three rounds of PCR amplification, and the flanking sequence could be obtained according to the sequence information obtained in the first step if the length obtained in a single experiment could not satisfy the experimental requirements.
This product is paired with a superior performance L-Exp Taq HS DNA Polymerase, with the ability to amplify long fragments or complex fragments with high GC content, is paired with a carefully optimized Buffer system to make the reaction highly specific, while at the same time obtaining longer target sequences with a guaranteed amplification success rate.
Control Template and Control Specific Primer are provided to facilitate control experiments.
The flanking sequences of the known sequences were obtained after three rounds of PCR amplification using the four concatenated primers provided in the kit, i.e., TP Primer 1 (TP1), TP Primer 2 (TP2), TP Primer 3 (TP3), and TP Primer 4 (TP4), respectively, and paired with the specific primers designed according to the known sequences, i.e., Specific Primer 1 (SP1), Specific Primer 2 (SP2), and Specific Primer 3 (SP3) for thermal asymmetric PCR amplification. Specific Primer 1 (SP1), Specific Primer 2 (SP2) and Specific Primer 3 (SP3) were used for thermal asymmetric PCR amplification, and the flanking sequences of the known sequences could be obtained after three rounds of PCR amplification, and the flanking sequence could be obtained according to the sequence information obtained in the first step if the length obtained in a single experiment could not satisfy the experimental requirements.
This product is paired with a superior performance L-Exp Taq HS DNA Polymerase, with the ability to amplify long fragments or complex fragments with high GC content, is paired with a carefully optimized Buffer system to make the reaction highly specific, while at the same time obtaining longer target sequences with a guaranteed amplification success rate.
Control Template and Control Specific Primer are provided to facilitate control experiments.
Product Advantages
1. This product utilizes L-Exp Taq HS DNA Polymerase has the advantages of high amplification efficiency and low mismatch rate; it adopts optimized PCR reaction system, which is very suitable for PCR amplification of long fragments and complex templates, making it suitable for chromosome stepping.
2. This product contains four parallel primers, at least one of which can be paired with a specific primer to obtain a flanking sequence after three rounds of PCR amplification, resulting in a high success rate.
2. This product contains four parallel primers, at least one of which can be paired with a specific primer to obtain a flanking sequence after three rounds of PCR amplification, resulting in a high success rate.
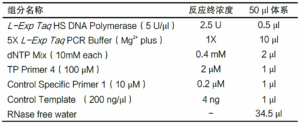
Product Composition
| individual parts making up a compound | norm |
| L-Exp Taq HS DNA Polymerase (5 U/µl) | 25 μl |
| 5X L-Exp Taq PCR Buffer (Mg)2+ plus) | 500 μl |
| dNTP Mix (10 mM each) | 100 μl |
| TP Primer 1 (100 µM) | 30 μl |
| TP Primer 2 (100 µM) | 30 μl |
| TP Primer 3 (100 µM) | 30 μl |
| TP Primer 4 (100 µM) | 60 μl |
| Control Template (200 ng / µl) | 10 μl |
| Control Specific Primer 1 (10 µM) | 10 μl |
| Control Specific Primer 2 (10 µM) | 10 μl |
| Control Specific Primer 3 (10 µM) | 10 μl |
Preservation and transportation
Storage temperature: -20℃ storage
Transportation temperature: dry ice transportation or -20℃ ice bag transportation
Transportation temperature: dry ice transportation or -20℃ ice bag transportation