This product is developed for Illumina high-throughput sequencing platform, based on the principle of CUT & RUN (Cleavage Under Targets and Release Using Nuclease) technology to study the interaction between protein and DNA. With optimized reaction system and library construction process, this product has the advantages of high experimental success rate, antibody compatibility, short experimental period, simple operation and high signal-to-noise ratio compared with traditional ChIP-Seq, which is especially suitable for epigenetic research fields such as early embryonic development, stem cells, tumorigenesis and so on.
CUT & RUN is a new method to study protein-DNA interaction, using Protein A/G fusion MNase nuclease to precisely target to the target protein under the guidance of antibody and fragmentation of DNA near the target site, the extracted DNA can be directly used for the construction of DNA libraries, and the DNA information of the target site can be obtained after sequencing.
The reaction system of this product has been carefully optimized, all reagents in the experiment should be used as provided in this product, it is not recommended to change any reaction components or replace the components in this product with other equivalent products in order to avoid obtaining bad experimental results. If replacement is needed, please verify it first.
1. For illumina sequencing platform: integrates Index and illumina connectors and constructs DNA libraries that can be directly sequenced by NGS.
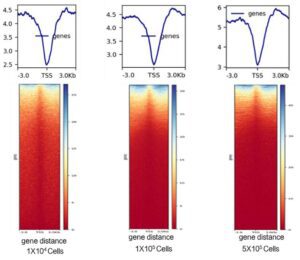
2. High compatibility: from 1 X 103 ~ 5 X 105 A starting volume of cells was experimented with.
3. High signal-to-noise ratio: low background noise signal, good experimental repeatability.
4. Simple operation: the whole process of magnetic bead extraction and purification, simple and rapid operation.
5. Strong performance of pAG-Mnase: pAG structural domain has broad-spectrum affinity for antibodies from rabbit, mouse and other sources, with precise targeting; optimized Mnase enzyme can efficiently and accurately cleave the target DNA.
6. High quality sequencing: optimized PCR amplification system, high quality sequencing data.
Package 2-1 is organized as follows (stored at 4°C):
| individual parts making up a compound | AG12562 | AG12561 |
| ConA Beads Pro | 20 μl | 60 μl |
| DNA Clean Beads | 470 μl | 1.5 ml |
| 10% SDS | 8 μl | 24 μl |
Package 2-2 is composed as follows (stored at -20°C):
| individual parts making up a compound | AG12562 | AG12561 |
| Lysis Buffer | 400 μl | 1.2 ml |
| 10X ConA Binding Buffer | 100 μl | 300 μl |
| 10X Wash Buffer | 700 μl | 1 ml X 2 pcs |
| 5% Digitonin Solution | 8 μl | 24 μl |
| 10X Dig-300 Buffer | 220 μl | 660 μl |
| BE Buffer | 14 μl | 42 μl |
| pAG-Mnase | 4 μl | 12 μl |
| CaCl2 Solution | 8 μl | 24 μl |
| Stop Buffer | 6 μl | 18 μl |
| Proteinase K (20 mg / ml) | 4 μl | 12 μl |
Package 2-2 -20℃ Storage
Transportation Temperature: Package 2-1 Ice Pack Transportation
Package 2-2 Dry Ice or -20°C Ice Pack Transportation